Banzai Japan
日本を元気に世界へ発信!47都道府県天下統一アイドル
もともと日本が持つ素晴らしい魅力である「山」「川」「建造物」「料理」「技術」「アニメ」「お・も・て・な・し」そして…「ヲタク」など!
日本がどや顔で誇れる文化を世界に向けて発信していくプロジェクトとして結成されたアイドルユニットです。
メンバーは、それぞれ日本の各都道府県を背負った伝道師として暗躍中だそうな。
47人揃うのは、いつの日や?
これから「BANZAI JAPAN(バンザイ ジャパン)」が、歌やダンスなどのパフォーマンスで、日本が世界に誇れるカルチャーを「見える化」して発信して参ります!
Japanese Recipes
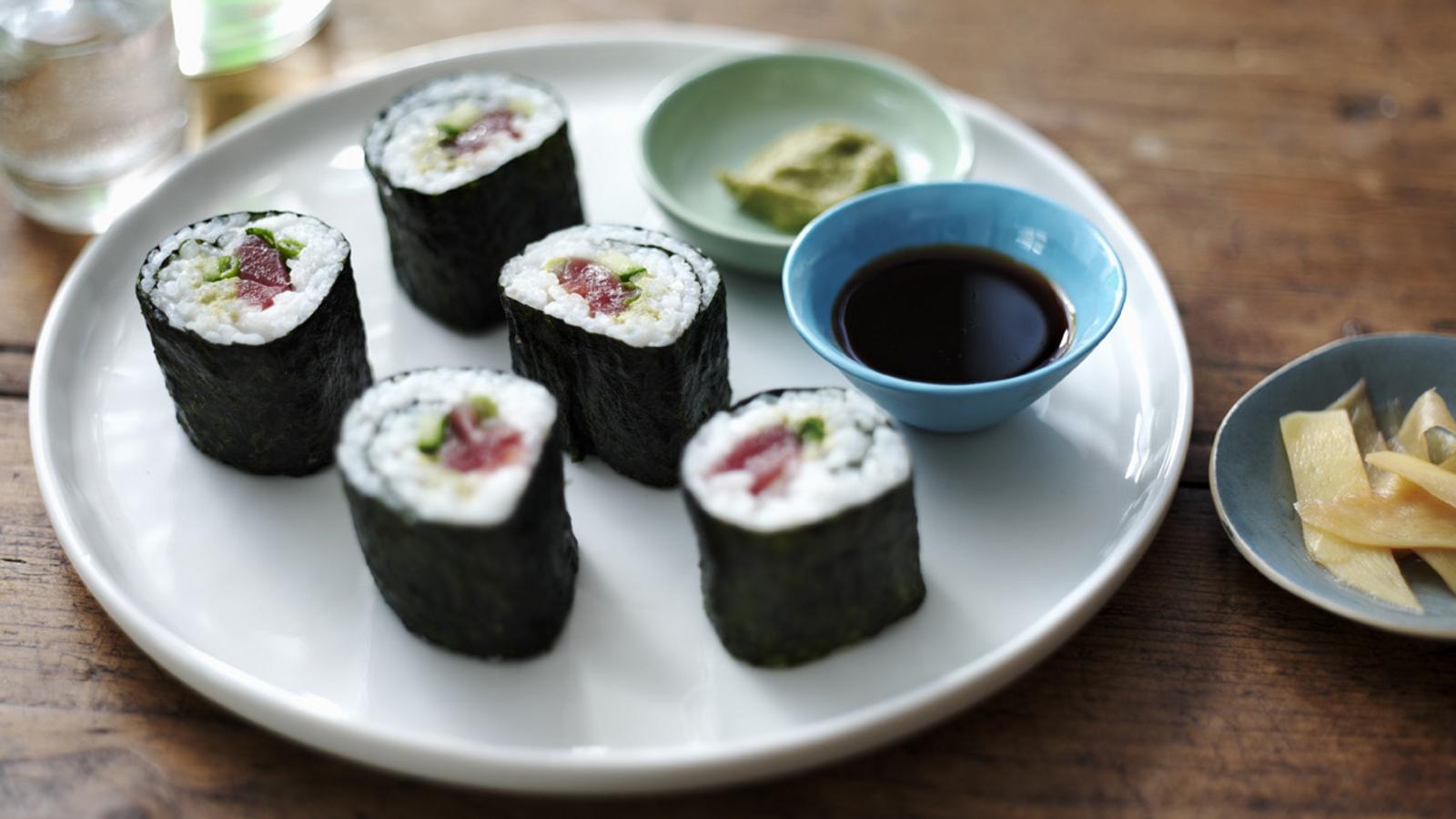

- Admin
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
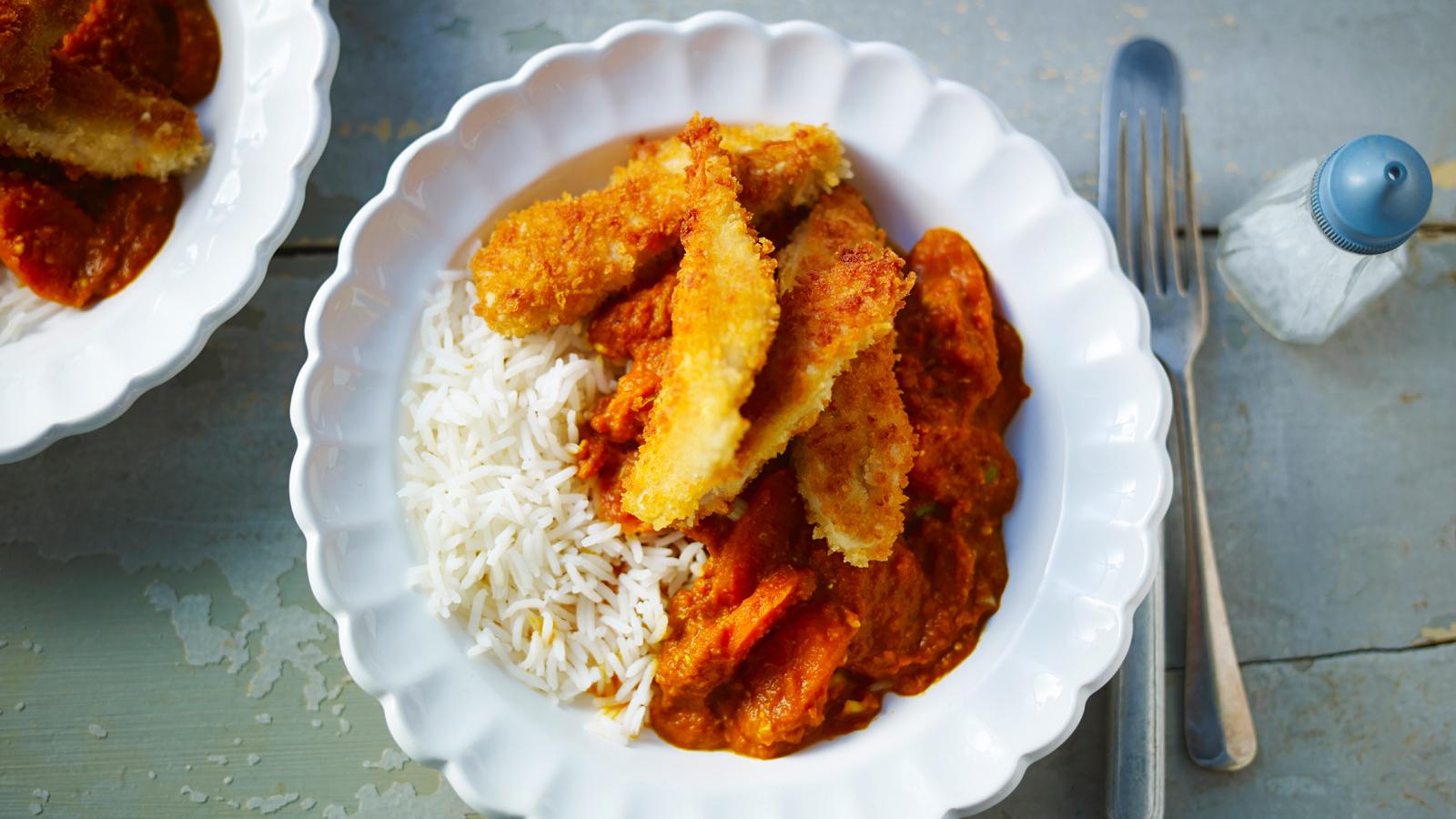

- Admin
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
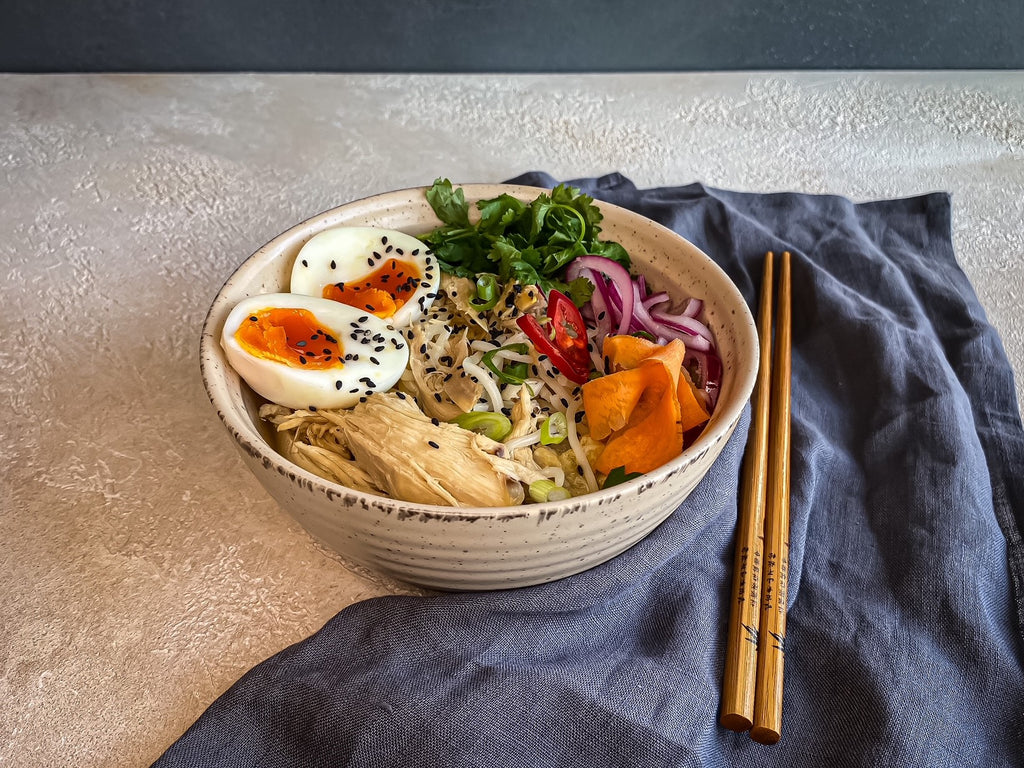

- Admin
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt

- Admin
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
British Recipes
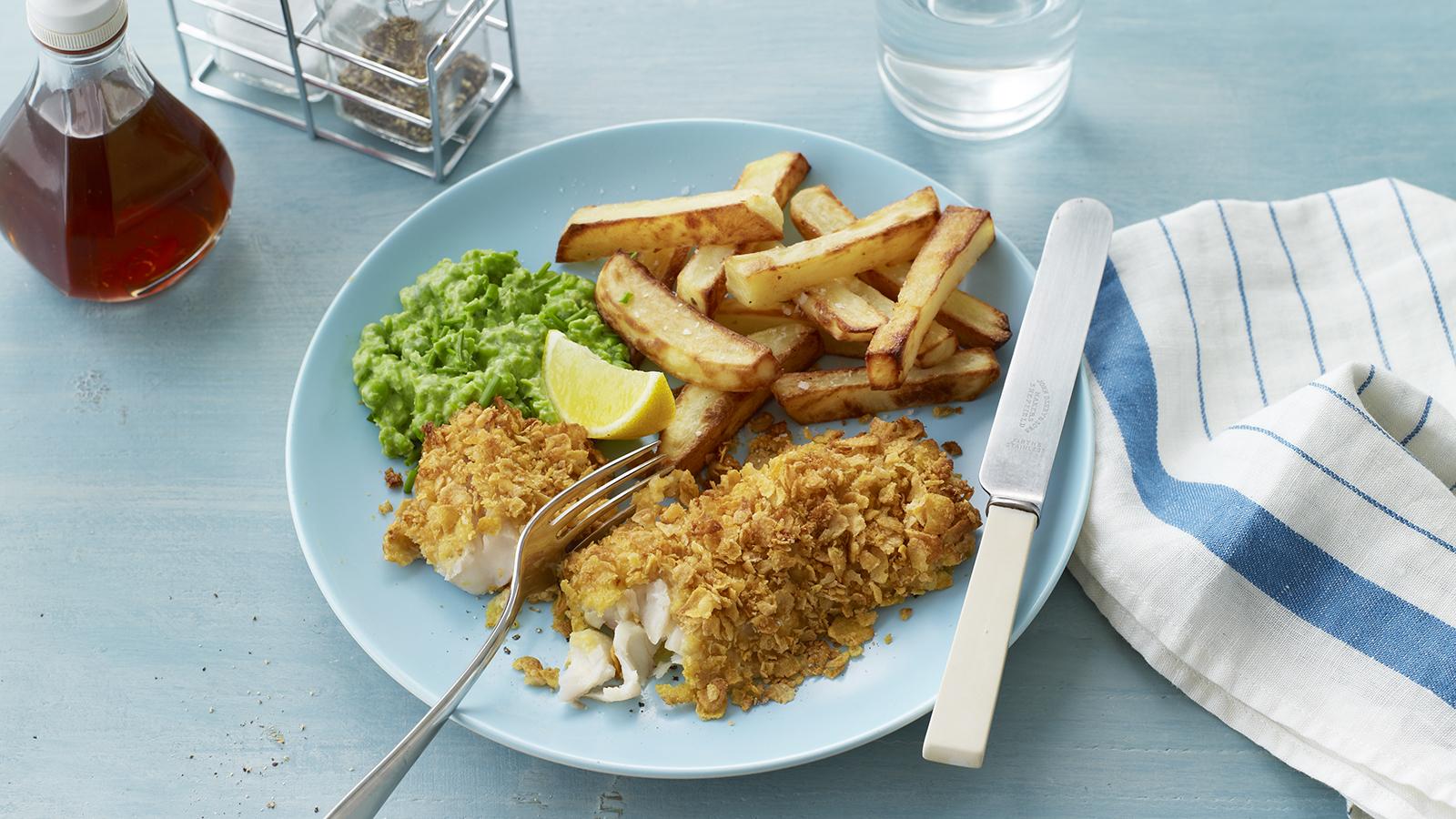

- Admin
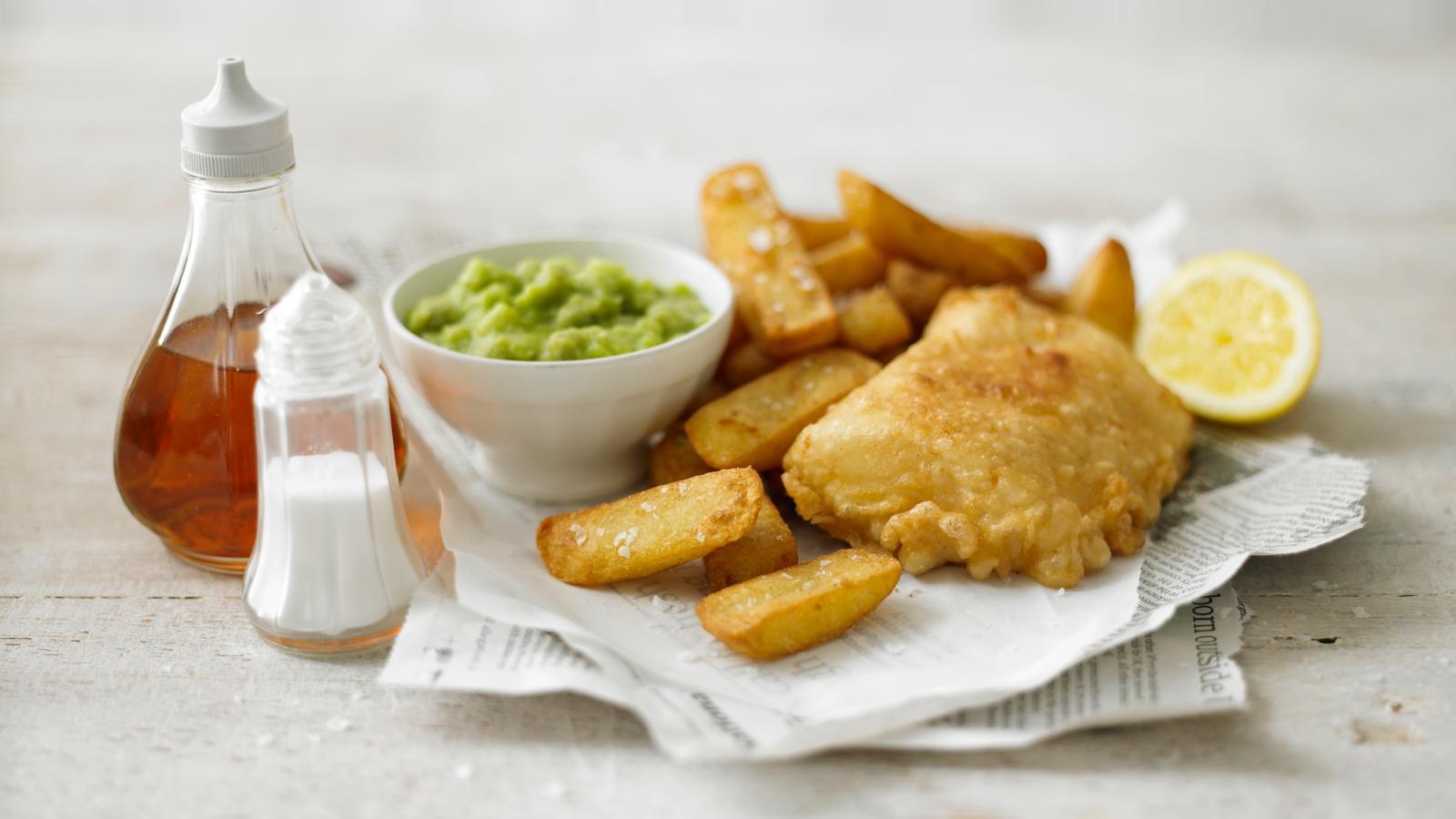
Eat Well for Less’ healthier and cheaper take on fish and chips. Cornflakes are the star of the show!
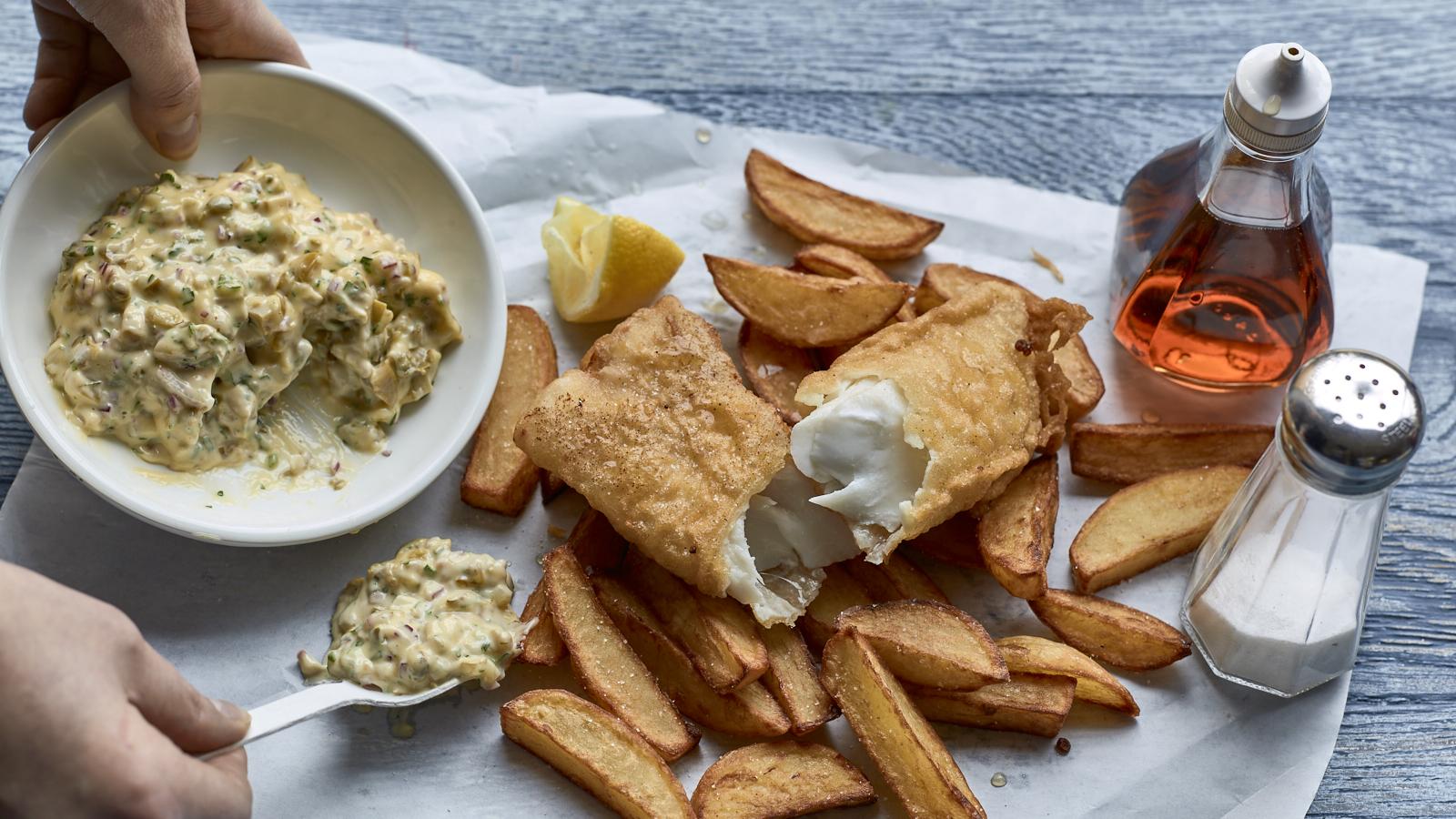

- Admin
A British classic of crunchy beer-battered fish and chips, with homemade tartare sauce. Serve with lots of salt and vinegar!

- Admin
Fish and chips with a few gourmet touches to make it special. The chip shop just won’t be the same after this.

- Admin
You can’t beat homemade fish & chips, get perfect results with Jo Pratt’s easy fish supper – a classic British favourite.
Featured
Popular Categories
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Ut elit tellus, luctus nec ullamcorper mattis, pulvinar dapibus leo.
Latest Recipes
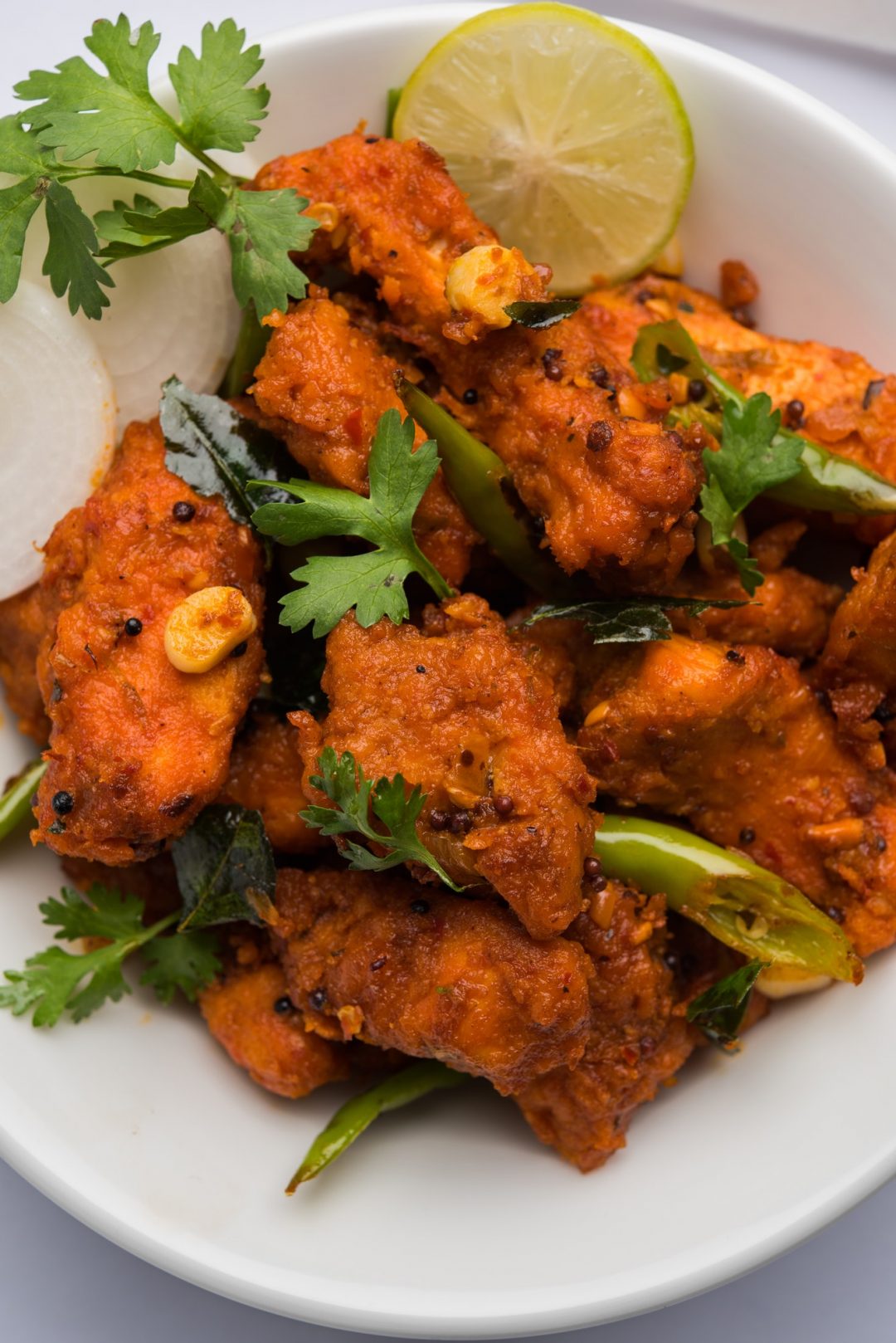

- by Bren
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
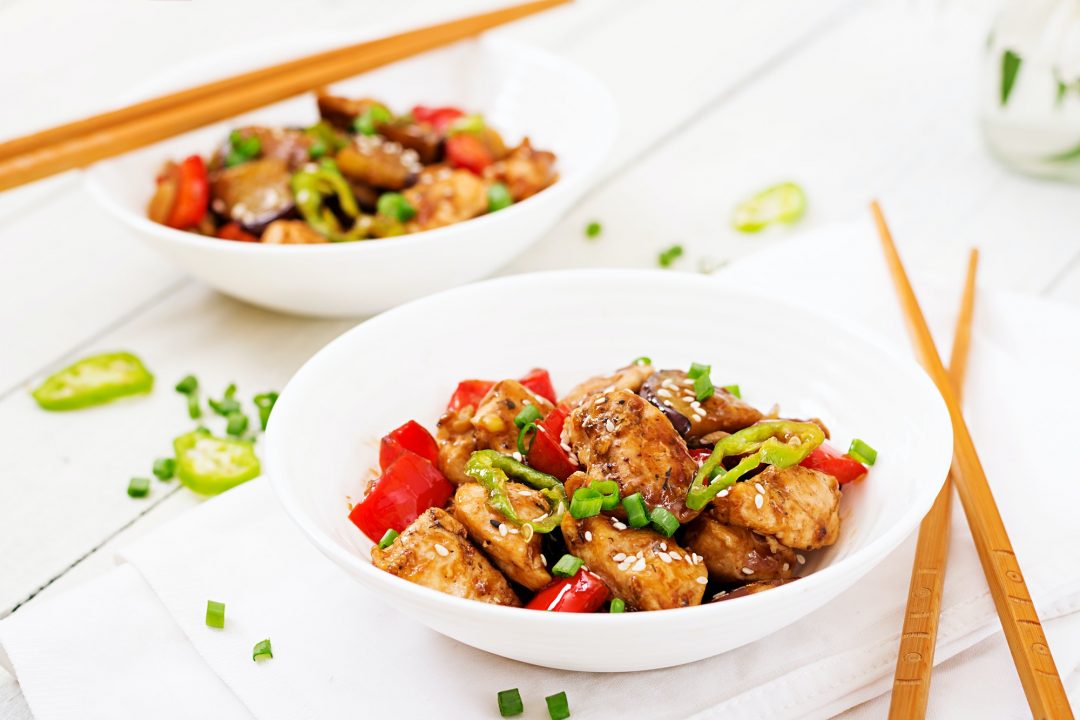

- by Morra
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
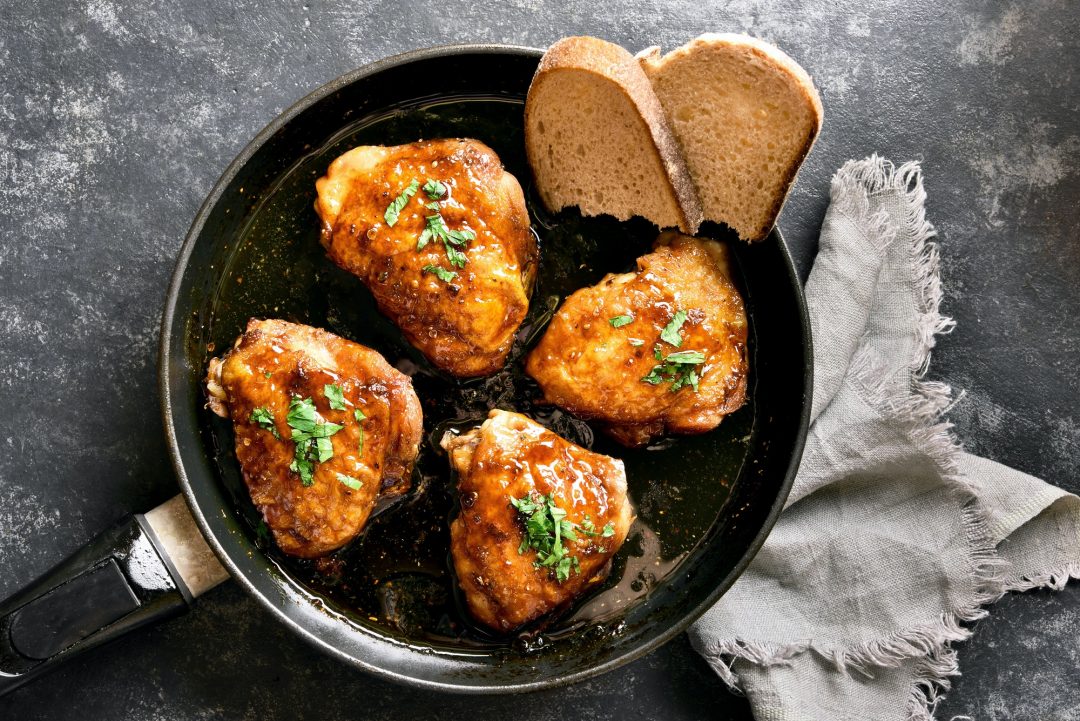

- by Chef Juna
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
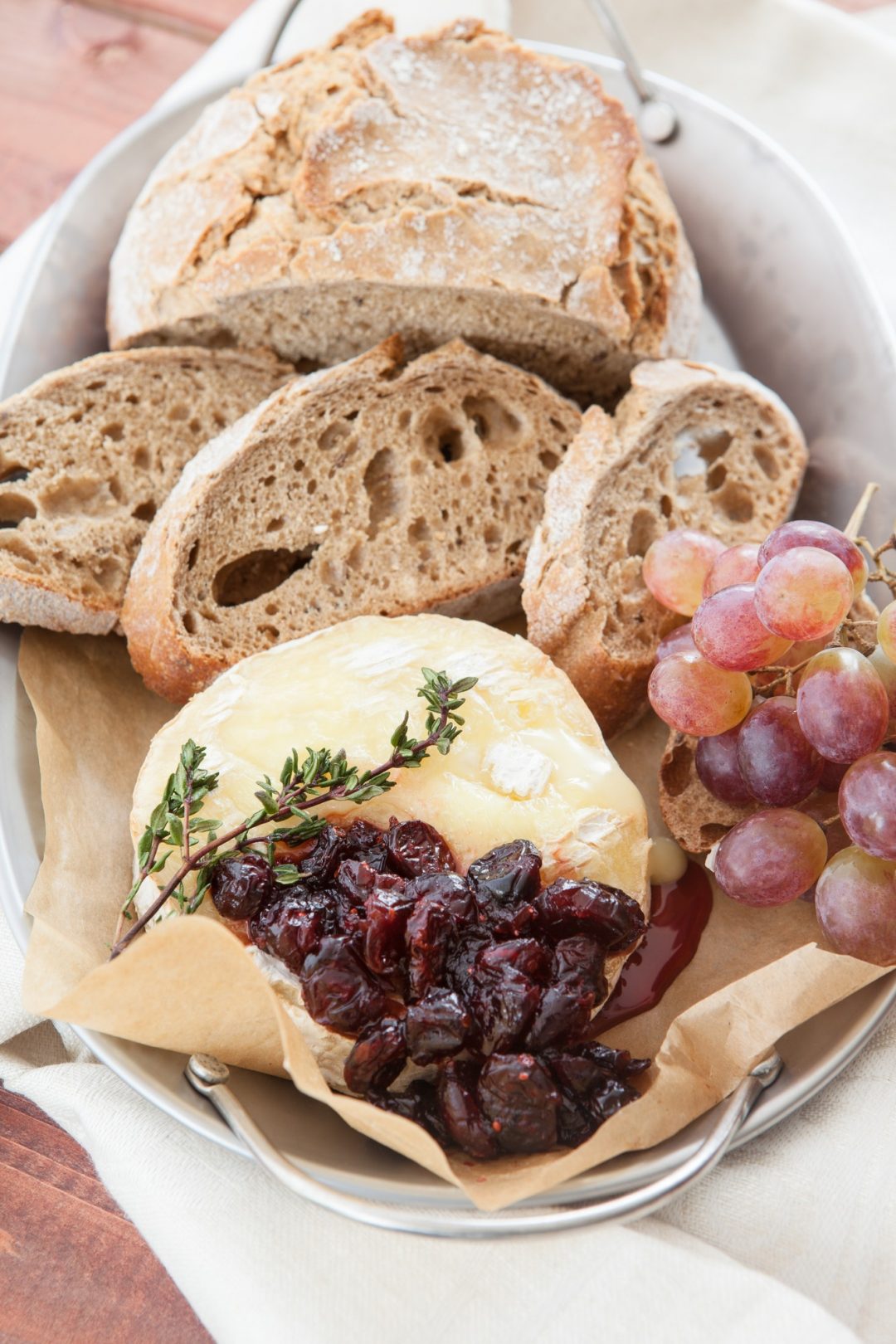

- by Wilgoz
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt

- by Bren
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
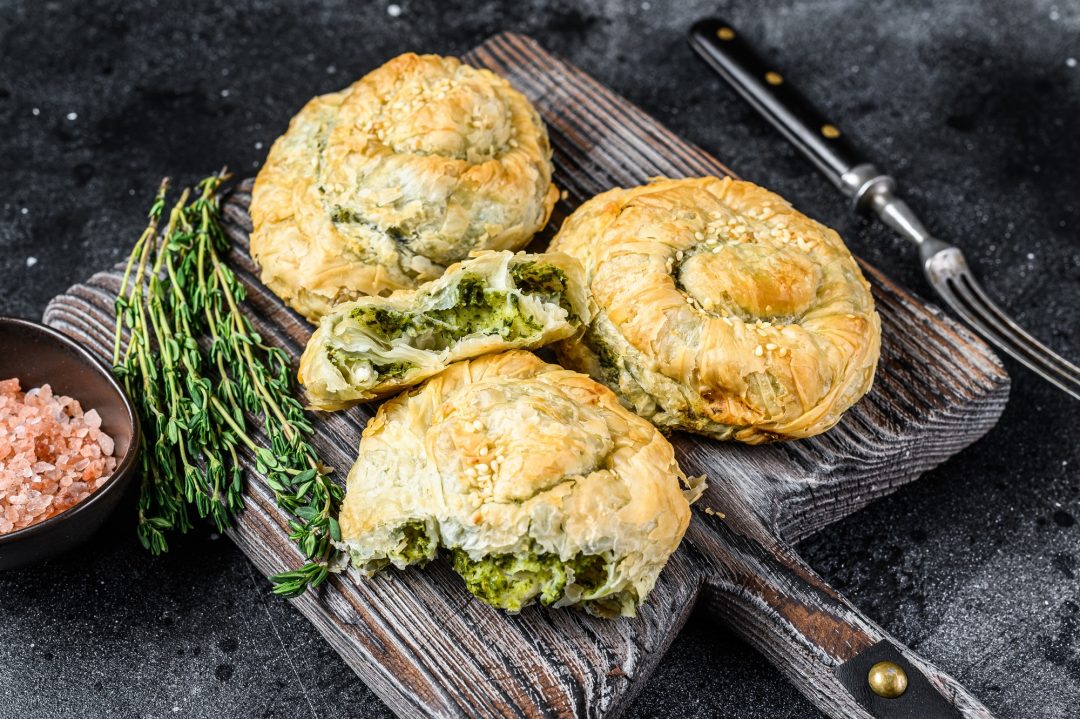

- by Morra
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
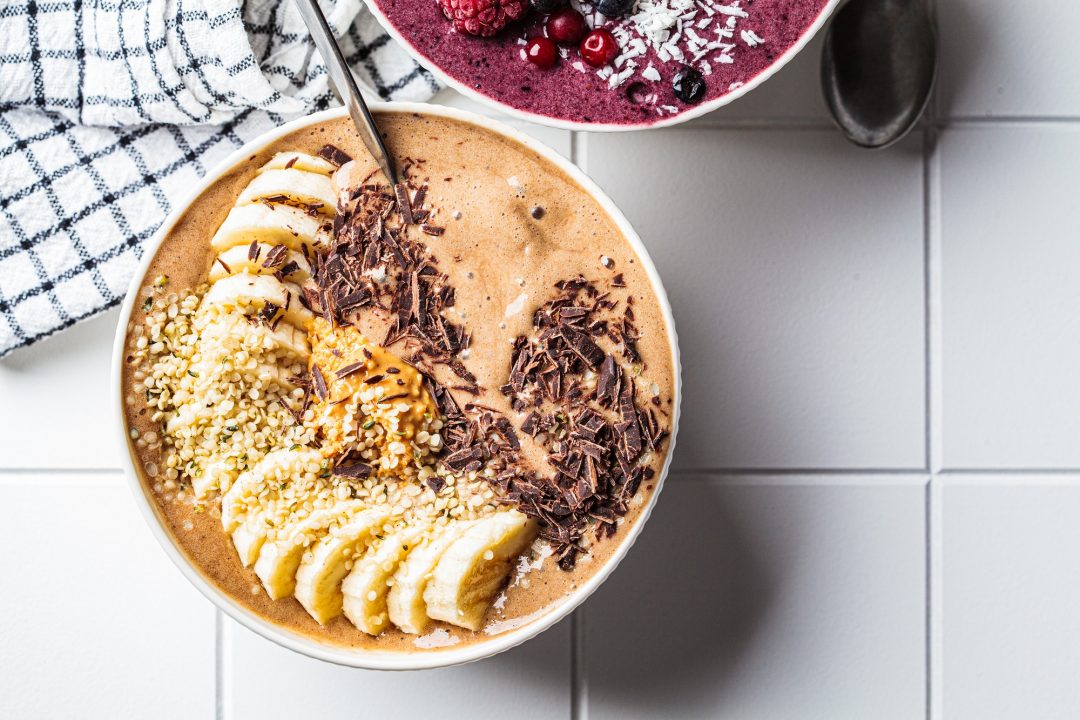

- by Chef Juna
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
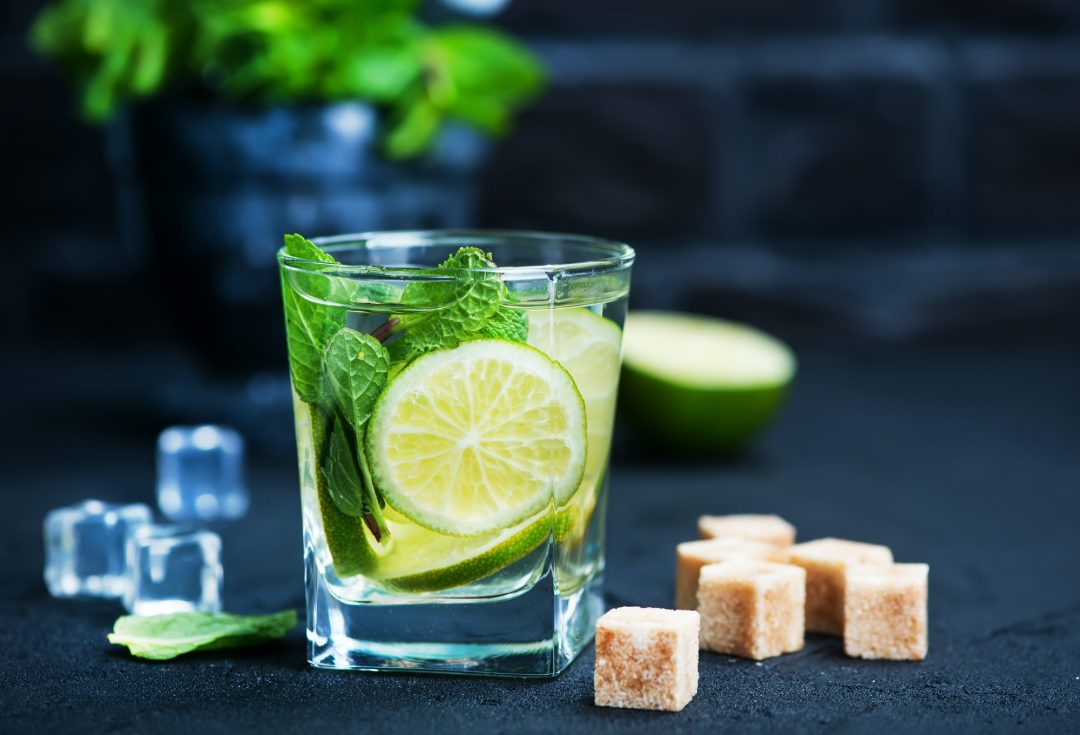

- by Wilgoz
Habitant accumsan suscipit sodales phasellus nulla elit placerat sapien quisque gravida tincidunt
Get update delicious recipes everyday!
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Ut elit tellus, luctus nec ullamcorper mattis, pulvinar dapibus leo.
Japan
Japan | History, Population, & Facts
Japan is an island country in East Asia. It is in the northwest Pacific Ocean and is bordered on the west by the Sea of Japan
Extending from the Sea of Okhotsk in the north toward the East China Sea, Philippine Sea, and Taiwan in the south.
Population
125.7 million (2021)
History
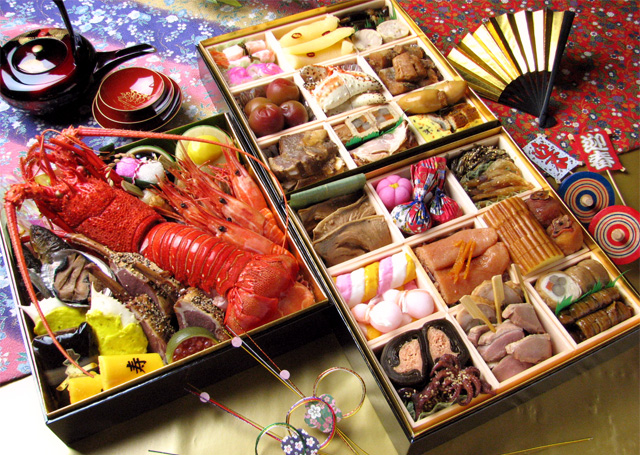
Japan is an archipelago, or string of islands, on the eastern edge of Asia. There are four main islands: Hokkaido, Honshu, Shikoku, and Kyushu. There are also nearly 4,000 smaller islands! Japan's nearest mainland neighbors are the Siberian region of Russia in the north and Korea and China farther south.
Japanese Fun Facts
1.It’s good manners to slurp your noodles 2.The traditional Christmas Eve meal is KFC 3.Japan is not all about the cities 4.There’s a Rabbit Island in Japan 5.The number four is extremely unlucky 6.There’s a bizarre naked festival 7.Japanese trains are some of the most punctual in the world 8.The Japanese love wacky flavours 9.Everyone has their own seal 10.Anti-Ninja floors are a thing


Testimonial
What they say
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Ut elit tellus, luctus nec ullamcorper mattis, pulvinar dapibus leo.



Message from founder
First, we eat. Then, we do everything else.
Consectetur quisque per vel ac aptent praesent convallis aliquam montes. Lectus sit inceptos dis dui mattis ad morbi leo luctus duis. Mauris neque dictumst letius consequat senectus potenti ad.
Consectetur quisque per vel ac aptent praesent convallis aliquam montes. Lectus sit inceptos dis dui mattis ad morbi leo luctus duis. Mauris neque dictumst letius consequat senectus potenti ad.